Hierarchical Vision Transformers for Prostate Biopsy Grading
Hierarchical Vision Transformers for Prostate Biopsy Grading
Towards Bridging the Generalization Gap
Deploying Vision Transformers on whole-slide images is non-trivial due to their extreme resolution. Drawing on Hierarchical Transformers developed for long documents, we adapt this paradigm to computational pathology and apply it to prostate cancer grading. We further introduce a novel attention factorization technique that combines attention scores across hierarchical levels, controlled by a single parameter γ that balances task-agnostic (pretrained) and task-specific (fine-tuned) contributions — enabling richer, more interpretable slide-level heatmaps.
Our best model achieves a quadratic kappa of 0.916 on the PANDA benchmark, matching state-of-the-art. Crucially, it generalizes better to diverse clinical settings, reaching a quadratic kappa of 0.877 on a crowdsourced multi-center dataset — outperforming all PANDA consortium teams evaluated on the same data.
Key Findings
Competitive in-distribution
Local H-ViT matches top PANDA challenge teams on same-center test data, confirming the hierarchical design does not sacrifice accuracy.
Best out-of-distribution generalization
On both the Karolinska external dataset and the more diverse Gleason grading in the Wild dataset, our model performs best overall, providing the strongest evidence of generalization to real-world clinical settings.
Interpretable attention factorization
A novel cross-hierarchical attention factorization method, controlled by a single parameter γ, balances task-agnostic pretraining features and task-specific fine-tuning signals.
Method
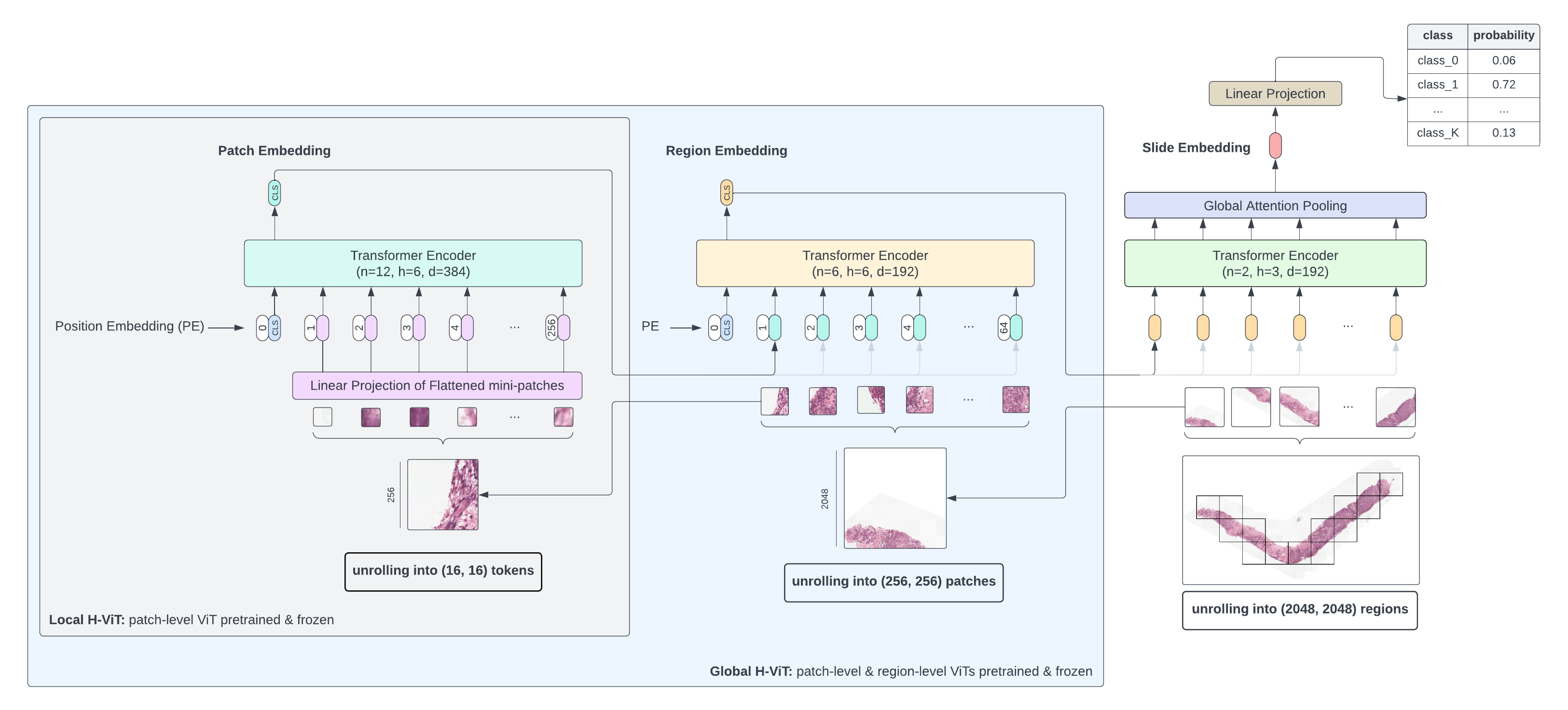
Whole-slide images exhibit a natural hierarchy of scales: from 16×16 pixel tokens capturing individual cell features, to 256×256 patches encoding cell-to-cell interactions, up to 2048×2048 regions capturing macro-scale tissue architecture.
H-ViT mirrors this structure with three stacked Transformers:
- Patch-level ViT — embeds 16×16 tokens within each 256×256 patch
- Region-level Transformer — aggregates patch embeddings into region representations
- Slide-level Transformer — pools region embeddings into a slide-level prediction
We explore two training configurations: Global H-ViT (only slide-level Transformer trained) and Local H-ViT (region- and slide-level Transformers trained jointly), and study the effect of region size, pretraining dataset, and loss function.
Attention Factorization
To improve interpretability, we introduce a unified heatmap that combines attention maps from all Transformer levels. For each pixel (x,y), the factorized attention score is:
\[a_{(x,y)} = \frac{1}{\beta} \sum_{i=0}^{N-1} a^i_{(x,y)} \left[\gamma(1 - \mathbf{1}_F(T_i)) + (1-\gamma)\mathbf{1}_F(T_i)\right]\]where γ ∈ [0,1] controls the balance between frozen (task-agnostic) and fine-tuned (task-specific) Transformers, and β is a normalization constant. For prostate grading, we recommend γ > 0.5 because it emphasizes coarser, task-specific features. This aligns with Gleason assessment, which is driven primarily by glandular growth patterns at the tissue-architecture scale rather than isolated cell-level cues.
Results
Comparison against PANDA consortium teams
Classification performance of our best ensemble Local H-ViT models against that PANDA consortium teams on PANDA public and private test sets, as well as Karolinska University Hospital dataset, used as external validation data after the challenge ended. All values are quadratic weighted kappa.
| Model | PANDA public | PANDA private | PANDA combined | Karolinska (external) |
|---|---|---|---|---|
Save The Prostate | 0.921 | 0.938 | 0.928 | 0.881 |
NS Pathology | 0.918 | 0.934 | 0.927 | 0.899 |
PND | 0.911 | 0.941 | 0.925 | 0.890 |
iafoss | 0.918 | 0.930 | 0.925 | 0.824 |
Aksell | 0.921 | 0.927 | 0.925 | 0.879 |
vanda | 0.922 | 0.930 | 0.922 | 0.880 |
BarelyBears | 0.912 | 0.933 | 0.920 | 0.890 |
| Local H-ViT (ours) | 0.915 | 0.917 | 0.916 | 0.895 |
Our model is competitive in-distribution and achieves the second best generalization on the Karolinska dataset.
The appearance of slides in the Karolinska University Hospital dataset is close to that of the PANDA dataset originating from the Karolinska Institute. This could explain why most models generalize well on this dataset and calls for additional evaluation in a more diverse clinical setting.
Real-World Stress Test: Generalization on a Multi-Center Dataset
Because of its crowdsourced nature, the prostate biopsy dataset introduced in Faryna et al. (2024) encompasses diverse clinical settings (tissue preparation protocol, staining, scanning device). This multi-center dataset uniquely represents the full diversity of cases encountered in clinical practice, allowing for a more rigorous assessment of model performance in a real-world setting.
| Model | QW Kappa (κ²) | Accuracy | Binary Accuracy |
|---|---|---|---|
PND | 0.862 | 0.513 | 0.876 |
BarelyBears | 0.845 | 0.531 | 0.903 |
NS Pathology | 0.760 | 0.611 | 0.876 |
Kiminya | 0.716 | 0.513 | 0.867 |
vanda | 0.617 | 0.336 | 0.779 |
| Local H-ViT (ours) | 0.877 | 0.602 | 0.903 |
Binary accuracy = distinguishing low-risk (ISUP ≤ 1) from higher-risk cases.
Overall, our model performs on par with or better than all compared models across the reported metrics on the crowdsourced dataset: while most PANDA challenge models show a marked performance drop, our best model maintains strong generalization.
BibTeX
@article{grisi2025,
title = {Hierarchical Vision Transformers for prostate biopsy grading:
Towards bridging the generalization gap},
author = {Clément Grisi and Kimmo Kartasalo and Martin Eklund and
Lars Egevad and Jeroen {van der Laak} and Geert Litjens},
journal = {Medical Image Analysis},
volume = {105},
pages = {103663},
year = {2025},
doi = {10.1016/j.media.2025.103663}
}